Dr. Li-Meng Yan presents evidence of the artificial origin of COVID-19. Now the protesters will be happy


The interview with Dr. Li-Meng Yan has warmed the spirits and some (even replying to us on twitter….) Have asked for proof of her statements. Well, the Chinese academician didn't wait long and, after opening a Twitter account, she started posting the scientific basis of her claims.
Clearly they are not understandable documents at all (maybe @Chimicoscettico could take a look at them …) but we try to provide understandable extracts.
First of all, he published a link to his scientific paper with a rather clear title:
As a reinforcement to the research, he has placed his academic profile which results in 13 scientific papers and his affiliation to the University of Hong Kong
Here are some excerpts:
Evidence shows that SARS-CoV-2 is expected to be a laboratory product created using the bat coronaviruses ZC45 and / or ZXC21 as a model and / or backbone. Based on the evidence, we further postulate a synthetic pathway for SARS-CoV-2, demonstrating that the laboratory creation of this coronavirus is cost-effective and can be accomplished in about six months.
Here is the extended punchline:
The reason is that the SARS-CoV-2 Spike receptor binding cannot arise from nature and should have been created through genetic engineering.
Spike proteins decorate the outside of the coronavirus particles. They play an important role in infection as they mediate interaction with host cell receptors and thus help determine the host range and tissue tropism of the virus. The Spike protein is divided into two halves (Figure 3). The anterior or N-terminal half is called S1, which is completely responsible for binding the host receptor. In both SARS-CoV and SARS-CoV-2 infections, the host cell receptor is hACE2. Within S1, a segment of approximately 70 amino acids comes into direct contact with hACE2 and is correspondingly referred to as the binding form of the receptor (RBM) (Figure 3C). In SARS-CoV and SARS-CoV-2, RBM fully determines the interaction with hACE2. The C-terminal half of the Spike protein is named S2. The primary function of S2 includes the maintenance of trimer formation and, following protease cleavages following the S1 / S2 junction and an S2 'position downstream, mediate fusion of the
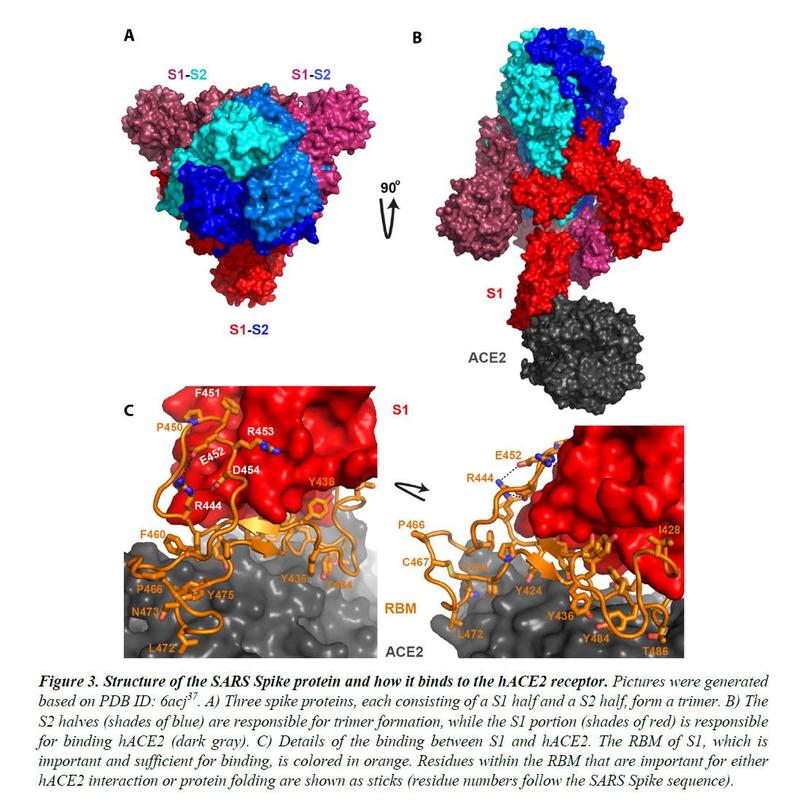
Similar to that observed for other viral proteins, S2 from SARS-CoV-2 shares a high sequence identity (95%) with S2 from ZC45 / ZXC21. In stark contrast, between SARS-CoV-2 and ZC45 / ZXC21, the S1 protein, which determines which host (human or bat) the virus can infect, is much less conserved with amino acid sequence identity by only 69%.
Figure 4 shows the sequence alignment of Spike proteins from six β coronaviruses. Two are viruses isolated from the current pandemic (Wuhan-Hu-1, 2019-nCoV_USA-AZ1); there are two suspected template viruses (Bat_CoV_ZC45, Bat_CoV_ZXC21); two are SARS coronaviruses (SARS_GZ02, SARS). The RBM is highlighted between two orange lines. Clearly, despite the high sequence identity for the overall genomes, the RBM of SARS-CoV-2 differs significantly from those of ZC45 and ZXC21. Curiously, SARS-CoV-2's RBM largely resembles SARS Spike's RBM. While this is not an exact "copy and paste", a careful examination of the Spike-hACE237,38 structures reveals that all residues essential for hACE2 binding or protein folding (orange sticks in Figure 3C and what is highlighted by short red lines in Figure 4) are “preserved”.
Most of these essential residues are precisely conserved, including those involved in disulfide bond formation (C467, C474) and electrostatic interactions (R444, E452, R453, D454), which are critical for the structural integrity of RBM. (Figure 3C and 4). The few changes within the group of essential residues are almost exclusively hydrophobic “substitutions” (I428àL, L443àF, F460àY, L472àF, Y484àQ), which should not affect either protein folding or interaction with hACE2. At the same time, most of the non-essential amino acid residues are “mutated” (Figure 4, unlabeled RBM residues with short red lines). Judging from the sequence analysis alone, we were convinced from the start that not only would the SARS-CoV-2 Spike protein bind to hACE2, but the binding would also be similar, precisely, to that between the original SARS Spike protein and hACE223. Recent structural work confirmed our prediction.
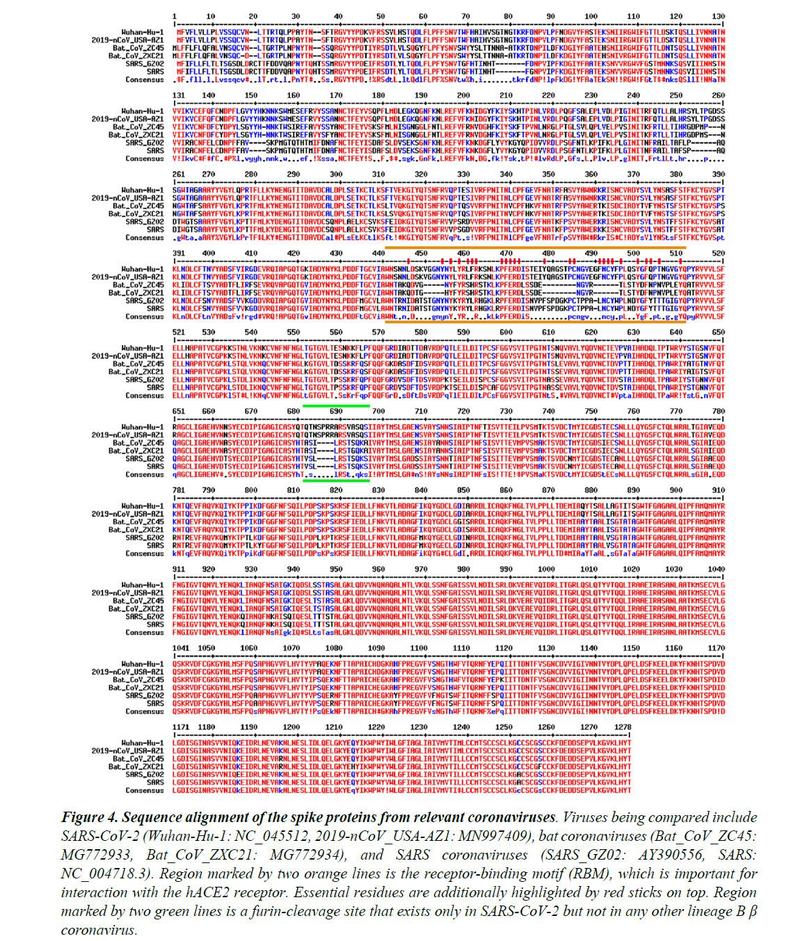
then the scientist makes two different hypotheses that may be at the basis of the "natural" birth of the virus. Whether this results from: 1) an ancient recombination event followed by convergent evolution or 2) a natural recombination event that occurred fairly recently.
She then goes on to exclude the two hypotheses:
1) “this process of convergent evolution would also involve the accumulation of a large amount of mutations in other parts of the genome, making the overall sequence identity relatively low. The high sequence identity between SARS-CoV-2 and ZC45 / ZXC21 on various proteins (94-100% identity) does not support this scenario and thus clearly indicates that SARS-CoV-2 carrying such an RBM it cannot come from a bat coronavirus similar to ZC45 / ZXC21 through this convergent evolutionary route ”.
Then delete option 2) 2) In the second scenario, the ZC45 / ZXC21-like coronavirus would have recently had to recombine and swap its RBM with another coronavirus that had successfully adapted to bind a highly homologous animal ACE2 to hACE2. The probability of such an event depends, in part, on the general requirements of natural recombination: 1) that the two different viruses share a significant sequence similarity; 2) which must co-infect and be present in the same cell of the same animal; 3) that the recombinant virus would not have been eliminated from the host or would not have caused the host to become extinct; 4) that the recombinant virus should eventually become stable and transmissible within the host species.
Regarding this recent recombination scenario, the animal reservoir could not be bats because the ACE2 proteins in bats are not homologous enough to hACE2 and therefore the adaptation would not be able to produce an RBM sequence as seen in SARS-CoV. -2. This animal reservoir also could not be human as the ZC45 / ZXC21-like coronavirus would not be able to infect humans. Additionally, there was no evidence of SARS-CoV-2 or SARS-CoV-2-like viruses circulating in the human population before the end of 2019. Curiously, according to a recent bioinformatics study, SARS-CoV-2 was well-adapted for humans since the beginning of the epidemic.
Discarded the impossible, what remains, however improbable, is the truth:
Only one more possibility of natural evolution remains, which is that the ZC45 / ZXC21-like virus and a coronavirus containing a SARS-like RBM may have recombined in an intermediate host where the ACE2 protein is homologous to hACE2. Several labs have reported that some of the Sunda pangolins smuggled into China from Malaysia were carriers of coronavirus, whose receptor-binding domain (RBD) is nearly identical to that of SARS-CoV-227-29,31. They then went on to suggest that pangolins are the likely intermediate host for SARS-CoV-227-29,31. However , recent independent reports have found significant flaws in these data . Furthermore, contrary to these reports, no coronaviruses were detected in Sunda pangolin samples collected for over a decade in Malaysia and Sabah between 2009 and 2019. A recent study also showed that RBD, which is shared between SARS-CoV -2 and the reported coronavirus pangolin, binds to hACE2 ten times stronger than pangolin ACE22, further excluding pangolins as a possible intermediate host. Finally, an in silico study, while echoing the idea that pangolins are likely not an intermediate host, also indicated that none of the animal ACE2 proteins examined in their study showed more favorable binding potential to the SARS-CoV protein. -2 Spike versus hACE2. This latest study exempted virtually all animals from their suspected intermediate host role, which is consistent with the observation that SARS-CoV-2 was well adapted for humans since the outbreak began. This is significant because these results collectively suggest that no intermediate host appears to exist for SARS-CoV-2, which at the very least decreases the possibility of a recombinant event occurring in an intermediate host.
Given that RBM fully enforces hACE2 binding and that SARS RBM-hACE2 binding was fully characterized by high resolution structures (Figure 3), this RBM-only exchange would be no more risky than the full Spike exchange. Indeed, the feasibility of this RBM trading strategy has been demonstrated. In 2008, Dr Zhengli Shi's team swapped a SARS RBM into the spike proteins of several SARS-like bat coronaviruses after introducing a restriction site into a codon-optimized spike gene (Figure 5C). They then validated the binding of the resulting chimeric Spike proteins with hACE2. Furthermore, in a recent publication, SARS-CoV-2 RBM was swapped into the receptor binding domain (RBD) of SARSCoV, resulting in a fully functional chimeric RBD in binding hACE2 (Figure 5C). Surprisingly, in both cases, the manipulated RBM segments almost exactly resemble the RBM defined by the locations of the EcoRI and BstEII sites (Figure 5C). Although cloning details are lacking in both publications39,47, it is conceivable that actual restriction sites may vary depending on the spike gene receiving RBM insertion, as well as the convenience of introducing unique restriction sites in regions of interest. Interestingly, the corresponding author of this recent publication, Dr. Fang Li, has been an active collaborator of Dr. Zhengli Shi since 2010. Dr. Li was the first person in the world to structurally elucidate the link between SARS -CoV RBD and hACE238 and was the leading expert in the structural understanding of Spike-ACE2 interactions. The surprising discovery of EcoRI and BstEII restriction sites at either end of the SARS-CoV-2 RBM, respectively, and the fact that the same RBM region was swapped by both Dr. Shi and his long-term collaborator, respectively, using restriction enzyme digestion methods are unlikely as a coincidence. Rather, it is the smoking gun that proves that SARS-CoV-2's RBM / Spike is a product of genetic manipulation. "
It seems that the Chinese also tried to hide the evidence …
While it may be convenient to copy the exact sequence of SARS's RBM, it would be too clear a sign of artificial design and manipulation. The most deceptive approach would be to modify some non-essential residues, while preserving those critical for the bond. This design could well be guided by high resolution structures (Figure 3) 37,38. Thus, when the overall sequence of RBM would appear to be more distinct from that of SARS RBM, the ability to bind hACE2 would be well preserved. We believe that all the crucial residues (residues labeled with red sticks in Figure 4, which are the same residues shown in the sticks in Figure 3C) should have been "preserved". As described above, while some should be direct storage, some should be switched to residues with similar properties, which do not break the binding to hACE2 and may even further strengthen the binding [ZH: i.e., the virus has been armed and boosted ]. Importantly, the changes may have been made intentionally to non-essential sites, making it less like a “copy and paste” of SARS RBM .
So there was no genetic manipulation, but the authors did everything they could to try to hide it. A truly embarrassing situation for the Chinese government if it were scientifically confirmed internationally and which would justify all the accusations made.

Thanks to our Telegram channel you can stay updated on the publication of new articles of Economic Scenarios.
The article Dr. Li-Meng Yan presents evidence of the artificial origin of COVID-19. Now the protesters will be happy comes from ScenariEconomici.it .
This is a machine translation of a post published on Scenari Economici at the URL https://scenarieconomici.it/la-dottoressa-li-meng-yan-presenta-le-prove-dellorigine-artificale-del-covid-19-ora-i-contestatori-saranno-contenti/ on Mon, 14 Sep 2020 18:00:51 +0000.
